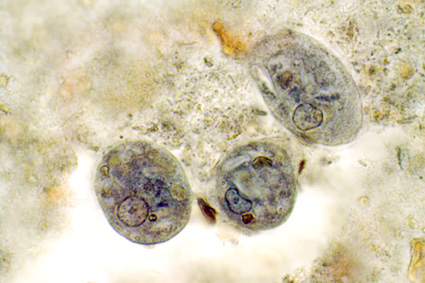
The month of September saw the publication of the first data on Blastocystis subtypes going out from Qatar. Abu-Madi and colleagues--who have already been quite prolific in terms of surveying intestinal parasitic infections in Qatar--studied the positive rate of Blastocystis in 608 apparently healthy subjects arriving in Qatar for the first time, identifying a prevalence of 71% as identified by PCR. Strikingly, the positive rate by microscopy of the corresponding samples was only 7%. Three subtypes were idenfied, with ST3 being the most common subtype, followed in prevalence by ST1 and ST2. The study is important for at least two reasons: It confirms the drawback of basing Blastocystis epidemiological research on data generated using microscopy alone, and it confirms the virtual absence of ST4 outside of Europe.
Increased sensitivity of PCR relative to microscopy was also confirmed in a study carried out in Malaysia (I presume) by Ragavan and colleagues. This group surveyed the Blastocystis positivity rate among IBS and non-IBS patients analyzing colonic aspirates, including a total of 109 individuals. Given the data available on Blastocystis prevalence, I was quite surprised to learn that this group failed to detect Blastocystis in any of the samples by microscopy and culture. Using PCR (the subtype-specific [STS] primers were used as diagnostic primers), the group identified Blastocystis in 6 IBS patients and 4 non-IBS patients. Also these figures appear quite low. However, there is very little information available on the non-IBS patients, and since all study individuals were subject to colonscopy, this group of individuals might be suffering chronic and potentially severe intestinal disease, including for instance colorectal cancer, inflammatory bowel disease, etc., which would explain the low prevalence of Blastocystis observed among these individuals. Indeed, evidence is accumulating that the more "gut healthy" you are, the larger the probability of being Blastocystis-positive. I noticed that the colonic aspirates were spun down using 3,000 rpm prior to culture and microscopy; this process might have had an impact on cell viability and morphology; still, DNA should be detectable following this process. Meanwhile, we recently showed (Scanlan et al., 2015) that the sensitivity of the STS primers is relatively low, which is why the use of real-time PCR is recommendable for PCR-based screening. To see an example of how the STS primers perform relative to barcoding primers, go here (Suppl Table 2).
Moreover, care should be taken when reading this paper, since I'm fairly convinced that the subtype terminology used in the study is different from the consensus terminology (Stensvold et al., 2007). It says that the subtypes detected included ST2, ST3, ST4, and ST5; if this reflects the terminology that went along with the original description of the STS primers, these subtypes correspond to ST7, ST3, ST6, and ST2, which to me would be a more likely subtype distribution, taking this particular region into consideration, and given the fact that ST5 appears to be extremely rare in humans.
It's always interesting to expand on the natural host spectrum of Blastocystis. The parasite has been found in a perplexing array of hosts, but some host specificity has been observed. When it comes to animals held by humans as livestock or pets, we know that pigs and cattle are commonly, if not consistently, colonised by Blastocystis with some quite specific subtypes. With regard to pets, dogs and cats have been found positive, but there seems to be increasing evidence that these animals are not natural hosts (see also Wang et al., 2013). Osman and colleagues, recently published a survey on Cryptosporidium and Blastocystis in dogs using sensitive molecular methods, demonstrating a prevalence of Blastocystis of only about 3%. Moreover, the subtypes 2 and 10 were found, and ST10 is found mostly in cattle, and never before in dogs, as far as I know, which could suggest accidental colonisation - and possibly not a very long-lasting one. Similarly, when humans are found to be colonised with subtypes rarely found in humans, such as ST6, ST7, and ST8, it would be interesting to know for how long these subtypes are capable of "staying put" in the human intestine.
References
Abu-Madi M, Aly M, Behnke JM, Clark CG, & Balkhy H (2015). The distribution of Blastocystis subtypes in isolates from Qatar. Parasites & Vectors, 8 PMID: 26384209
Increased sensitivity of PCR relative to microscopy was also confirmed in a study carried out in Malaysia (I presume) by Ragavan and colleagues. This group surveyed the Blastocystis positivity rate among IBS and non-IBS patients analyzing colonic aspirates, including a total of 109 individuals. Given the data available on Blastocystis prevalence, I was quite surprised to learn that this group failed to detect Blastocystis in any of the samples by microscopy and culture. Using PCR (the subtype-specific [STS] primers were used as diagnostic primers), the group identified Blastocystis in 6 IBS patients and 4 non-IBS patients. Also these figures appear quite low. However, there is very little information available on the non-IBS patients, and since all study individuals were subject to colonscopy, this group of individuals might be suffering chronic and potentially severe intestinal disease, including for instance colorectal cancer, inflammatory bowel disease, etc., which would explain the low prevalence of Blastocystis observed among these individuals. Indeed, evidence is accumulating that the more "gut healthy" you are, the larger the probability of being Blastocystis-positive. I noticed that the colonic aspirates were spun down using 3,000 rpm prior to culture and microscopy; this process might have had an impact on cell viability and morphology; still, DNA should be detectable following this process. Meanwhile, we recently showed (Scanlan et al., 2015) that the sensitivity of the STS primers is relatively low, which is why the use of real-time PCR is recommendable for PCR-based screening. To see an example of how the STS primers perform relative to barcoding primers, go here (Suppl Table 2).
Moreover, care should be taken when reading this paper, since I'm fairly convinced that the subtype terminology used in the study is different from the consensus terminology (Stensvold et al., 2007). It says that the subtypes detected included ST2, ST3, ST4, and ST5; if this reflects the terminology that went along with the original description of the STS primers, these subtypes correspond to ST7, ST3, ST6, and ST2, which to me would be a more likely subtype distribution, taking this particular region into consideration, and given the fact that ST5 appears to be extremely rare in humans.
It's always interesting to expand on the natural host spectrum of Blastocystis. The parasite has been found in a perplexing array of hosts, but some host specificity has been observed. When it comes to animals held by humans as livestock or pets, we know that pigs and cattle are commonly, if not consistently, colonised by Blastocystis with some quite specific subtypes. With regard to pets, dogs and cats have been found positive, but there seems to be increasing evidence that these animals are not natural hosts (see also Wang et al., 2013). Osman and colleagues, recently published a survey on Cryptosporidium and Blastocystis in dogs using sensitive molecular methods, demonstrating a prevalence of Blastocystis of only about 3%. Moreover, the subtypes 2 and 10 were found, and ST10 is found mostly in cattle, and never before in dogs, as far as I know, which could suggest accidental colonisation - and possibly not a very long-lasting one. Similarly, when humans are found to be colonised with subtypes rarely found in humans, such as ST6, ST7, and ST8, it would be interesting to know for how long these subtypes are capable of "staying put" in the human intestine.
References
Abu-Madi M, Aly M, Behnke JM, Clark CG, & Balkhy H (2015). The distribution of Blastocystis subtypes in isolates from Qatar. Parasites & Vectors, 8 PMID: 26384209
Osman M, Bories J, El Safadi D, Poirel MT, Gantois N, Benamrouz-Vanneste S, Delhaes L, Hugonnard M, Certad G, Zenner L, & Viscogliosi E (2015). Prevalence and genetic diversity of the intestinal parasites Blastocystis sp. and Cryptosporidium spp. in household dogs in France and evaluation of zoonotic transmission risk. Veterinary Parasitology PMID: 26395822
Ragavan, N., Kumar, S., Chye, T., Mahadeva, S., & Shiaw-Hooi, H. (2015). Blastocystis sp. in Irritable Bowel Syndrome (IBS) - Detection in Stool Aspirates during Colonoscopy PLOS ONE, 10 (9) DOI: 10.1371/journal.pone.0121173
Scanlan PD, Stensvold CR, & Cotter PD (2015). Development and Application of a Blastocystis Subtype-Specific PCR Assay Reveals that Mixed-Subtype Infections Are Common in a Healthy Human Population. Applied and Environmental Microbiology, 81 (12), 4071-6 PMID: 25841010
Stensvold CR, Suresh GK, Tan KS, Thompson RC, Traub RJ, Viscogliosi E, Yoshikawa H, & Clark CG (2007). Terminology for Blastocystis subtypes--a consensus. Trends in Parasitology, 23 (3), 93-6 PMID: 17241816
Wang W, Cuttell L, Bielefeldt-Ohmann H, Inpankaew T, Owen H, & Traub RJ (2013). Diversity of Blastocystis subtypes in dogs in different geographical settings. Parasites & vectors, 6 PMID: 23883734